Boltz-2 predictions of protein-ligand structures and binding affinities with Python / Google Colab
Автор: PracticalCheminformatics
Загружено: 2025-10-07
Просмотров: 1110
Описание:
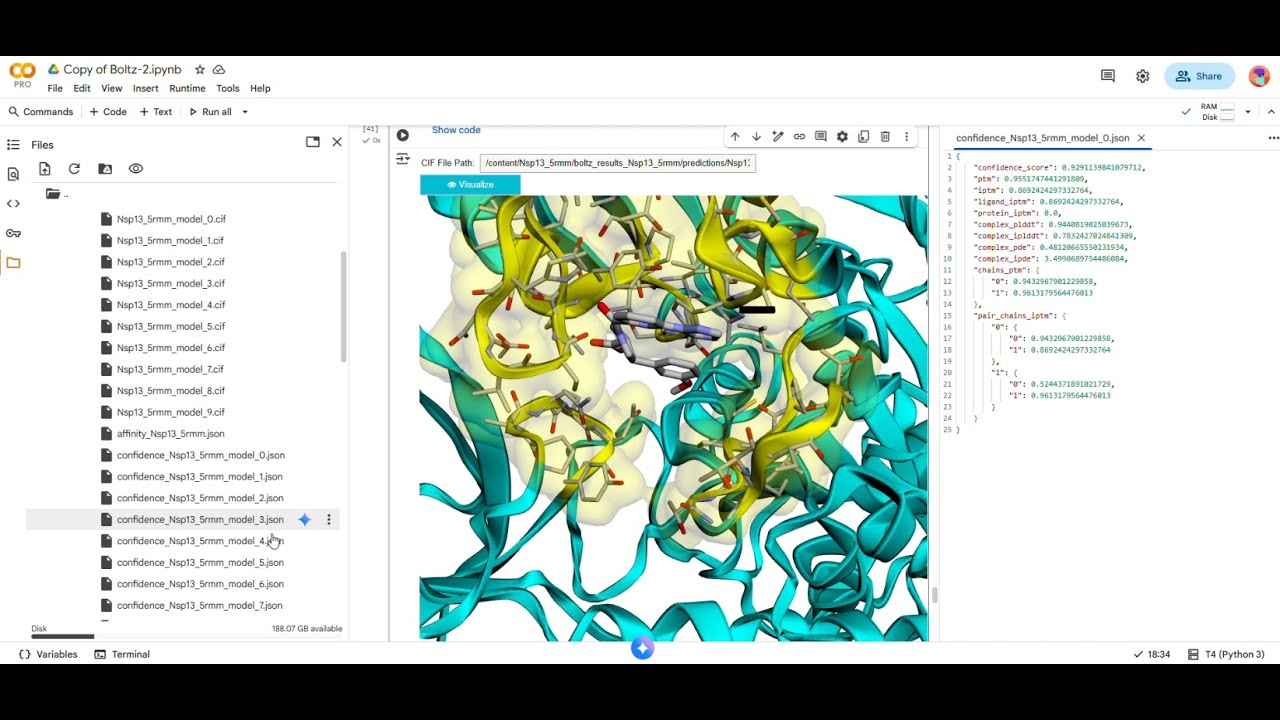
In this video, we walk through the use of Boltz-2 using python and Google Colab. This includes ligand design from an existing fragment so that structures and binding poses can be directly compared.
Boltz-2 is a generative AI model for structure‑based drug design that predicts protein–ligand complexes and associated binding affinities much faster than traditional physics‑based methods. [references: https://www.biorxiv.org/content/10.11... , https://github.com/jwohlwend/boltz]
Notably, Boltz-2 claims to be the first AI model to approach the performance of free-energy perturbation (FEP) methods in estimating small molecule–protein binding affinity while being at least 1000× more computationally efficient than FEP.
While full assessment of Boltz-2's capabilities will require extensive benchmarking and external verification, scientists have already started using Boltz-2 for their own drug discovery projects. A useful discussion on use of Boltz-2 on a range of different targets is discussed here; https://www.deepmirror.ai/post/boltz-...
In this example, I found the binding pose for the grown fragment to be in close agreement with an existing crystallographic pose, however the experimental binding affinity remains undetermined, so a direct comparison of theoretical vs experimental is not known.
Повторяем попытку...
Доступные форматы для скачивания:
Скачать видео
-
Информация по загрузке: