Automated genetic fine-mappingof neurological disorders
Автор: Neurogenomics Lab @ Imperial College London
Загружено: 2023-12-23
Просмотров: 34
Описание:
Summary
At the London Genetics Network Meeting (https://www.londongeneticsnetwork.com/) 2020, Brian Schilder ( / brian-schilder ) presents research on statistical inference of causal genomic variants underlying neurodegenerative conditions (Parkinson's Disease, Alzheimer's Disease, ALS, etc.), and the cell type-specific mechanisms by which they mediate their effects. This work was performed in collaboration with his colleagues Jack Humphrey ( / jack-humphrey-9ba9a26a ) and Towfique Raj ( / towfique-raj-8120836 ) @IcahnSchoolofMedicine
Background
When one conducts a GWAS or QTL study, we end up with a set of regions in the genome which contain variants associated with the given disease or phenotype. However, non-independence between variants due to linkage disequilibrium (LD) makes it very difficult to distinguish between the causal variant (or variants) underlying the disease, and variants that are simply highly correlated with the causal variants.
Fine-mapping is a class of statistical methods that seek to resolve these issues of causality, but is often underutilized. This is largely due to the technical difficulty of setting up idiosyncratic input/output formats across different programming languages, gathering the necessary data to run these tools (such as LD panels or genome-wide functional annotations). Also, each fine-mapping tools has its own set of strength and weakness, and can produce different (but often overlapping) credible sets of variants with 95% probability of being causal.
We therefore developed an R package called echolocatoR (https://github.com/RajLabMSSM/echoloc...) to integrate many of the most commonly used tools and datasets to seamlessly perform fine-mapping across all loci. All that is required by the user is a path to the full GWAS summary stats file and certain column names.
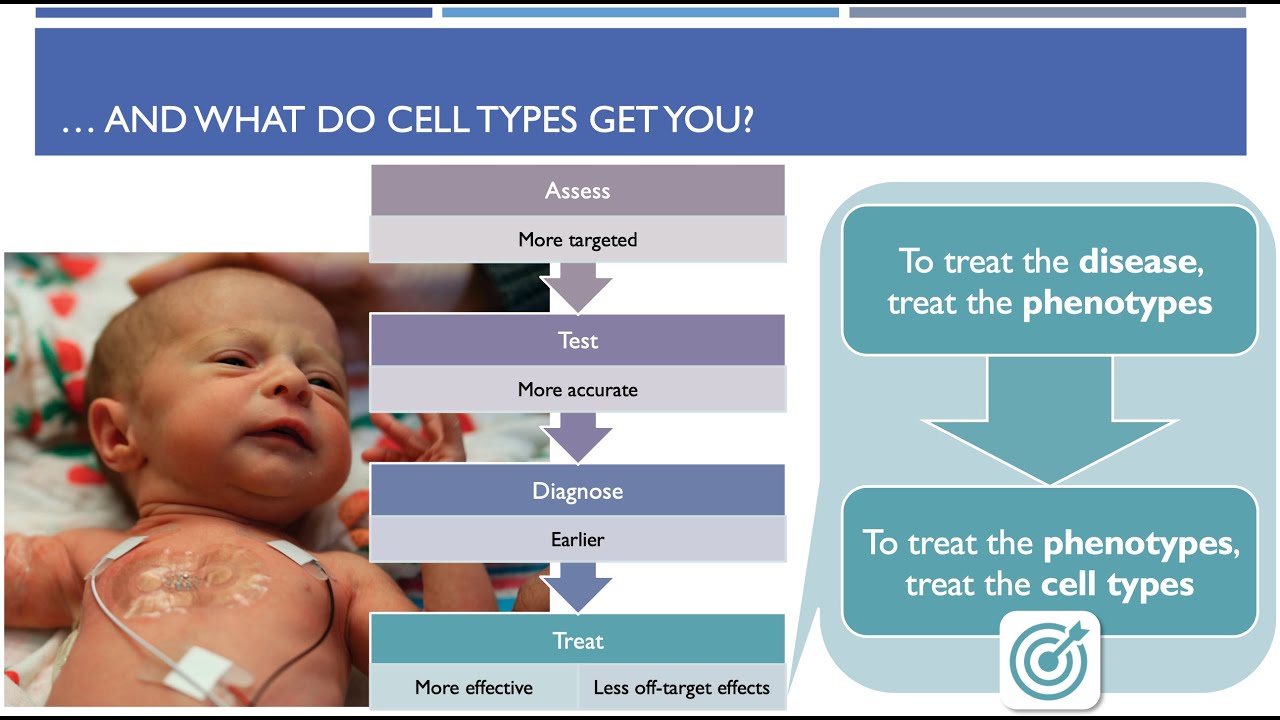
To conclude, echolocatoR can help to not only dissect genetic and cellular architecture of a given disease as a whole, like Parkinson’s, but also to make predictions about the primary mechanisms at play within a given individual’s particular form of the disease, in an effort to move to a more personalized approach to disease diagnosis and treatment.
Linked Resources
echolocatoR GitHub repository (https://github.com/RajLabMSSM/echoloc...)
Raj Lab (https://rajlab.org/index.html) @IcahnSchoolofMedicine
Linked Publications
echolocatoR: an automated end-to-end statistical and functional genomic fine-mapping pipeline (https://doi.org/10.1093/bioinformatic...)
Fine-mapping of Parkinson’s disease susceptibility loci identifies putative causal variants (https://doi.org/10.1093/hmg/ddab294)
Multi-omic insights into Parkinson's Disease: From genetic associations to functional mechanisms (https://doi.org/10.1016/j.nbd.2021.10...)
Genetic analysis of the human microglial transcriptome across brain regions, aging and disease pathologies (https://doi.org/10.1038/s41588-021-00...)
Dysregulation of mitochondrial and proteolysosomal genes in Parkinson’s disease myeloid cells (https://doi.org/10.1038%2Fs43587-021-...)
Повторяем попытку...

Доступные форматы для скачивания:
Скачать видео
-
Информация по загрузке: