iMeta: Applications of de Bruijn graphs in microbiome research
Автор: iMeta Science
Загружено: 2022-03-10
Просмотров: 1000
Описание:
Keith Dufault-Thompson, Xiaofang Jiang. 2022. Applications of de Bruijn graphs in microbiome research. iMeta 1: e4. https://doi.org/10.1002/imt2.4
Abstract
High-throughput sequencing has become an increasingly central component of microbiome research. The development of de Bruijn graph-based methods for assembling high-throughput sequencing data has been an important part of the broader adoption of sequencing as part of biological studies. Recent advances in the construction and representation of de Bruijn graphs have led to new approaches that utilize the de Bruijn graph data structure to aid in different biological analyses. One type of application of these methods has been in alternative approaches to the assembly of sequencing data like gene-targeted assembly, where only gene sequences are assembled out of larger metagenomes, and differential assembly, where sequences that are differentially present between two samples are assembled. de Bruijn graphs have also been applied for comparative genomics where they can be used to represent large sets of multiple genomes or metagenomes where structural features in the graphs can be used to identify variants, indels, and homologous regions in sequences. These de Bruijn graph-based representations of sequencing data have even begun to be applied to whole sequencing databases for large-scale searches and experiment discovery. de Bruijn graphs have played a central role in how high-throughput sequencing data is worked with, and the rapid development of new tools that rely on these data structures suggests that they will continue to play an important role in biology in the future.
Highlights
de Bruijn graph-based sequence assembly approaches have been an essential part of the broad application of sequencing methods, especially in microbiome research.
de Bruijn graphs can be used to efficiently represent sequencing data in a format that is highly scalable and can be extended and modified to address different research questions.
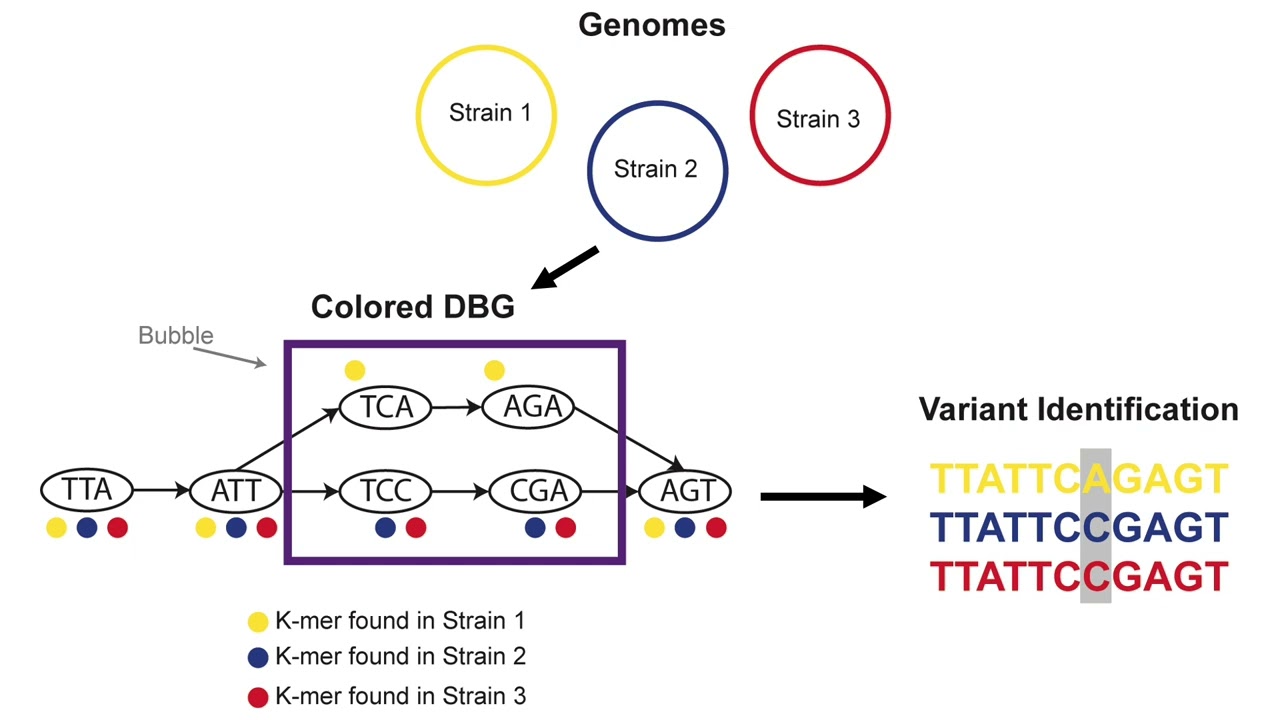
de Bruijn graph-based analysis methods have been developed for comparative genomics, the identification of genetic variants, and for large-scale searching of unassembled sequencing data.
The de Bruijn graph data structure will continue to be a central component of sequence assembly and analysis approaches in the future.
Повторяем попытку...
Доступные форматы для скачивания:
Скачать видео
-
Информация по загрузке: